Difference between revisions of "I-TASSER"
Jump to navigation
Jump to search
| Line 9: | Line 9: | ||
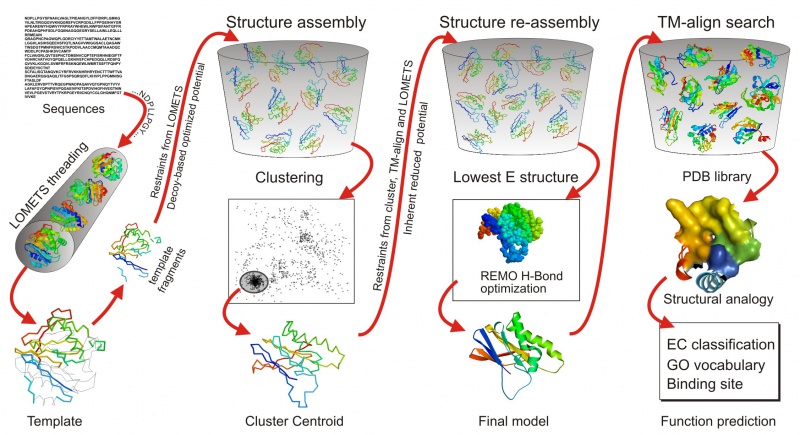
[[File:I-TASSER-pipeline.jpg|800px]] | [[File:I-TASSER-pipeline.jpg|800px]] | ||
| − | [https://zhanglab.ccmb.med.umich.edu/I-TASSER/example/ I-TASSER result | + | [https://zhanglab.ccmb.med.umich.edu/I-TASSER/example/ I-TASSER result example] |
===References=== | ===References=== | ||
[https://www.nature.com/articles/nprot.2010.5 '''I-TASSER: a unified platform for automated protein structure and function prediction''' (2010), Ambrish Roy, Alper Kucukural & Yang Zhang, Nature Protocols Volume 5, pages 725–738] | [https://www.nature.com/articles/nprot.2010.5 '''I-TASSER: a unified platform for automated protein structure and function prediction''' (2010), Ambrish Roy, Alper Kucukural & Yang Zhang, Nature Protocols Volume 5, pages 725–738] | ||
Revision as of 14:09, 13 June 2019
Welcome the I-TASSER tutorial page, a part of the Monmouth College Wiki.
The official link to the I-TASSER website can be found here.
The Wikipedia link can be found here.
- I-TASSER (Iterative Threading ASSEmbly Refinement) is a bioinformatics method for predicting three-dimensional structure model of protein molecules from amino acid sequences.[1] It detects structure templates from the Protein Data Bank by a technique called fold recognition (or threading). The full-length structure models are constructed by reassembling structural fragments from threading templates using replica exchange Monte Carlo simulations. I-TASSER is one of the most successful protein structure prediction methods in the community-wide CASP experiments.